Services
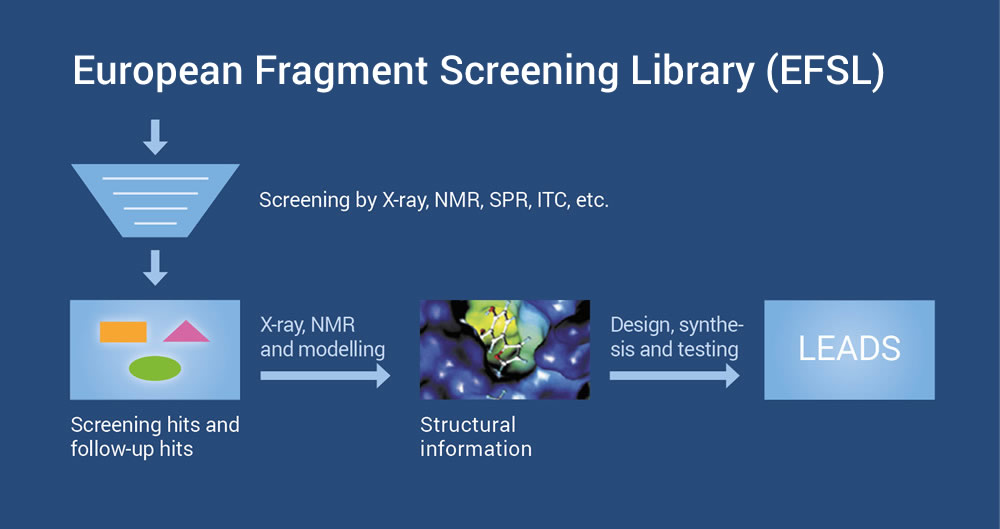
European Fragment Screening Library (EFSL)
EU-OPENSCREEN fragment-based lead discovery
EU-OPENSCREEN collaborates with the structural biology community (Instruct-ERIC/ iNEXT-Discovery) to offer fragment screening support, thereby exploiting their expertise in X-Ray crystallography, NMR spectroscopy, cryo-Electron Microscopy, among other techniques.
Instruct-ERIC/ iNEXT-Discovery and EU-OPENSCREEN partners joined efforts in biophysical chemistry, structural biology and medicinal chemistry to build a complete FBLDD pipeline, starting from the design of a novel fragment library, to executing fragment screening campaigns with hit identification all the way to evolution of lead compounds.
Keywords: lead identification, fragment screening, fragment evolution, fragment linking and fragment optimization
Please contact fragment-screening@eu-openscreen.eu for specific requests.
EU-OPENSCREEN Fragment Screening Library
The new EU-OPENSCREEN fragment screening library is composed of a set of low-molecular-weight fragments and ultra-low-molecular-weight so-called ‘minifrags’, derived from the fragment space of the EU-OPENSCREEN high-throughput diversity screening library. The fragment library has been so designed in order to facilitate the user follow-up with a rapid selection of parent molecules in EU-OPENSCREEN high-throughput diversity screening library, of which the fragments and minifrags are substructures.
The Fragment library is composed of:
- 968 fragments (available as 100 mM solutions in DMSO-d6 in 96-, 384-, and echo-compatible 1536-well plates)
- 88 minifrags (available as 1 M solutions in DMSO-d6 in 96- and 384-well plates).
Fragments were quality controlled by LC-MS and data can be downloaded here soon.
How to use the EU-OPENSCREEN Fragment Library
External researchers can use the Fragment Library in collaboration with EU-OPENSCREEN and Instruct-ERIC/ iNEXT-Discovery partners, following their respective application procedures.
The EU-OPENSCREEN fragment library is screened in external user projects as well as in in-house activities at the relevant partner sites. Before the start of each screening project, users (e.g. EU-OPENSCREEN or iNEXT-Discovery partners and/or their external users) have to complete the EU-OPENSCREEN user application form providing preliminary project information (contact details, project title and short description, duration of the screening, technology used etc), and send it back to fragment-screening@eu-openscreen.eu.
For more information on how to apply with a screening project please contact the partner of interest (partners information is available below), while for general enquiries please contact the EU-OPENSCREEN central office at fragment-screening@eu-openscreen.eu
For EU-OPENSCREEN and iNEXT-Discovery partners only: internal guidelines on the use of the EU-OPENSCREEN fragment library are available here (tba soon).
Where to screen the EU-OPENSCREEN Fragment Library
External researchers can use the Fragment Library in collaboration with EU-OPENSCREEN and Instruct-ERIC/ iNEXT-Discovery partners, following their respective application procedures. For more information, please contact us at fragment-screening@eu-openscreen.eu
EU-OPENSCREEN Fragment Screening partners
National Institute of Research and Innovation (NIRI)
Technology: Surface Plasmon Resonance(SPR), Ligand-based NMR spectroscopy, Isothermal Titration Calorimetry (ITC), and X-ray crystallography.
Contact Person: Aigars Jirgensons - aigars@osi.lv
Príncipe Felipe Research Center (CIPF)
Health Research Institute Hospital La Fe (IIS La Fe)
Technology: Surface Plasmon Resonance (SPR), Ligand-based NMR spectroscopy, and Isothermal Titration Calorimetry (ITC) including thepossibility of biochemical experiments, using different platforms, for validating the in-vitro activity of the fragments.
Contact: Antonio Pineda Lucena - pineda_ant@gva.es
University of Bergen (UiB)
Technology: Surface Plasmon Resonance (SPR)and Biolayer interferometry (BLI)
Contact: Ruth Brenk - ruth.brenk@uib.no
Natural Science Research Center of the Hungarian Academy of Sciences (MTA TTK) – an EU-OPENSCREEN-DRIVE partner
Technology: Fluorescence Polarization (FP), and self-developed kinase activity assays (luciferase enzyme complementation).
Contact Person: Attila Remenyi - remenyi.attila@ttk.mta.hu
iNext-Discovery partners
EMBL Grenoble - High-Throughput Crystallisation Laboratory (EU-OPENSCREEN-DRIVE site)
Technology: X-ray crystallography
Contact: José Márquez - marquez@embl.fr
Diamond Light Source - XChem facility
Technology: X-ray crystallography
Contact: Frank von Delft - frank.von-delft@diamond.ac.uk
Helmholtz Zentrum Berlin - Macromolecular crystallography at the BESSY-II electron storage ring
Technology: X-ray crystallography
Contact: Manfred Weiss - manfred.weiss@helmholtz-berlin.de
This site is screening a subselection of the fragment library. To compress the EU OPENSCREEN fragment library for pilot or lower throughput screenings, a 96-membered subselection was created. After applying some filters (e.g. 8-17 HA, practical filters like predicted solubility and minor manual selection) the remaining compounds were clustered by fingerprint similarities to arrive at a representative, yet sufficiently diverse subset of the entire EU OPENSCREEN fragment library. Please request details at fragment-screening@eu-openscreen.eu
Goethe University Frankfurt- Center for Biomolecular Magnetic Resonance (BMRZ)
Technology: Ligand-based NMR spectroscopy
Contact: Harald Schwalbe - Schwalbe@em.uni-frankfurt.de
FragMAX facility at MAX IV Laboratory
Technology: X-ray crystallography (please add any biophysical method if available)
Contact: Tobias Krojer tobias.krojer@maxiv.lu.se
What happens after the fragment screen?
After the fragment screen, users can access the EU-OPENSCREEN consultation support for the selection of appropriate fragment hits for chemical evolution and screen follow-up.
Moreover, The EU-OPENSCREEN project management team will collect the feedback on the screening campaign for the purpose of monitoring the performance and improving the EU-OPENSCREEN fragments library.
Please note that users of the EU-OPENSCREEN fragment library will be asked to make information on resulting hits publicly available in the European Chemical Biology Database (ECBD) and/ or other open-access databases such as PDBe (for structural data not supported by the ECBD). An embargo period of up to 36 months can be requested during the first 6 months after the first notification of a validated hit, to allow for publication and/ or filing patent applications to secure potential intellectual property. An extension of the embargo period after 36 months can be requested to the EU-OPENSCREEN office upon providing a written justification at fragment-screening@eu-openscreen.eu. Data will be automatically uploaded into the public domain after 6 months passed without an embargo requested.
Chemical consultation support for hit evolution
EU-OPENSCREEN offers chemical consultation support for hit evolution. The following medicinal chemists are renowned experts in chemical optimization of fragments into leads/ tool compounds: https://drive.eu-openscreen.eu/drive-startseite/about/medchem-consultants.html
Please find below a list of EU-OPENSCREEN chemical consultants:
Consultants at DTU (Denmark):
Name: Mads Hartvig Clausen and Faranak Nami.
Expertise: Fragment library synthesis (fluorinated fragments, diverse and shapely libraries), medicinal chemistry (hit-to-lead, in vitro ADME) and screening (ligand-observed NMR, thermal shift assay, FP).
Keywords: Fragment design and synthesis, NMR Screening, hit-to-lead chemistry.
Consultants at CSIC (Spain):
Name: Ana Martinez
Expertise: Small heterocyclic molecules, drug discovery and development of new drugs for the treatment of neurological diseases (Alzheimer’s, Parkinson’s, ALS, Multiple Sclerosis, etc.), molecular modelling and chemoinformatics, organic synthesis, biological screening, ADMETox properties optimization and technology transfer.
Keywords: Heterocyclic molecules, CNS, BBB, protein kinase inhibitors, small chemical probes, neurodegenerative diseases, neurological disorders
Name: Carmen Gil
Expertise: smallheterocyclic molecules. drug design and development, molecular modelling and chemoinformatics, organic synthesis, biological screening, ADMETox properties optimization and business management. research focuses on the search of new drugs for infectious diseases using both target-based and phenotypic approaches.
Keywords: Drug design, heterocyclic molecules, infectious diseases, protein kinase inhibitors, phosphodiesterase inhibitors, protein-protein interaction, ADME properties
Consultants at FMP (Germany):
Name: Marc Nazare
Expertise: Chemical probe discovery and development by screening approaches or structure-based design for selective targeting and cellular imaging. Hit-triage, chemical optimization of small molecule modulators of target classes such as GPCRs, kinases, phosphatases, proteases and ion channels. Structure-activity-relationships (SAR) and molecular recognition phenomena in protein-ligand interactions. Iterative multiparameter optimization of activity, selectivity, physico-chemical and ADME properties. Synthetic methodologies towards novel heterocyclic scaffolds useful in library design and probe discovery. FBDD-related expertise: Fragment based inhibitor evolution by fragment growing, merging and linking including multivalent approaches. Analysis of inhibitor binding by fragment deconstruction. Scaffold hopping and scaffold hybridization.
Keywords: FBDD, fragment growing, fragment merging, fragment linking, Scaffold hopping and scaffold hybridization, Hit-triage, SAR and chemical optimization of small molecule modulators.
Consultants at OSI (Latvia):
Name: Aigars Jirgensons
Expertise: Synthesis methodology and drug discovery, FBLD, chemical evolution of fragments by applying diverse synthetic modifications for growth and linking of fragment hits and structure-based lead optimization
Keywords: Serine protease, aspartic protease, cysteine protease, kinase, and PLP dependent enzyme inhibitors; GPCR, NMDA, and AMPA receptor ligands, hit-to-lead development, lead optimization; anti-bacterial, anti-malarial, CNS, anti-cancer drug discovery.
To select an EU-OPENSCREEN consultant, please fill in the Consultant Selection Form.
After acceptance of the consultation support for hit evolution, our medchem consultant will prepare a Fragment evolution report within 20 working days.
EU-OPENSCREEN consultants will propose follow-ups from the European Chemical Biology Library (ECBL) and commercially available fragment hit analogues. Moreover, they will provide a proposal of synthetic fragment follow-ups based on the compounds synthetic feasibility and propose fragment merging or fragment growth analogues, if structural information is available.
For EU-OPENSCREEN consultants only: the fragment evolution report template can be downloaded here.
Medicinal Chemistry support
Read more details on our medicinal chemistry services here.
For iNext Discovery and EU-OPENSCREEN-DRIVE partners only:
How to receive the fragment library
If you are an EU-OPENSCREEN-DRIVE and/ or iNEXT-Discovery partner who would like to receive the fragment library please fill in this Expression of Interest Form and send it back to fragment-screening@eu-openscreen.eu. The EU-OPENSCREEN management team will evaluate your request.
For iNEXT-Discovery sites: in case of eligibility, a collaboration agreement will be signed between the respective iNEXT-Discovery site and EU-OPENSCREEN ERIC with the purpose to formalise and regulate the terms and conditions for using and sharing of the EU-OPENSCREEN fragment library by and with the iNEXT-partner. The collaboration agreement template can be found here soon.
Acknowledgements
The fragment library was fully funded by the EU under grant agreement. No 823893 (EU-OPENSCREEN-DRIVE) and supplied by Specs (Fragment Library aggregation provider).